To all Apache Beam and Dataflow users:
If you’ve experimented with Beam, prototyped a pipeline, or verified assumptions about a dataset, you might have used Beam Notebooks or other interactive alternatives such as Google Colab or Jupyter Notebooks.
You might also have noticed a gap between running a small prototype pipeline in a notebook and a production pipeline on Dataflow: What if you want to interactively process and inspect aggregations of bigger production datasets from within the notebook, but at scale? You cannot rely on the single machine that’s running your notebook to execute the pipeline because it simply lacks the capacity to do so.
Allow me to introduce Interactive FlinkRunner on notebook-managed clusters. It lets you execute pipelines at scale and inspect results interactively with FlinkRunner on notebook-managed clusters. Under the hood, it uses Dataproc with its Flink and Docker components to provision long-lasting clusters.
This post will introduce you to Interactive FlinkRunner using three examples:
A starter word count example with a small notebook-managed cluster.
An example using a much bigger cluster to process tens of millions of flight records to see how many flights are delayed for each airline.
An example reusing the bigger cluster to run ML inference against 50,000 images with a pre-trained model – all from within a notebook.
If you want to control the cost1 of these examples, you are free to use pipeline options to reduce the size of the data and the cluster.
Prerequisites
Once you have Beam Notebooks instantiated, create an empty notebook (ipynb) file and open it with a notebook kernel selected2.
To get started, you have to check whether your project has the necessary services activated and permissions granted. You can find relevant information about the current user by executing the following in the notebook.
Interactive Flink on notebook-managed clusters uses Dataproc under the hood.
A starter example – Word Count
You’ve probably already seen the word count example multiple times. You know how to process and inspect the counted words with an InteractiveRunner or a DirectRunner on a single machine.
And you are able to run the pipeline on Dataflow as a one-shot job from within the exact same notebook without copying/pasting, moving across workspaces, or setting up the Cloud SDK.
To run it interactively with Flink on a notebook-managed cluster, you only need to change the runner and optionally modify some pipeline options.
The notebook-managed Flink cluster is configurable through pipeline options. You need these imports for this and the other examples.
You can then set up the configurations for development and execution.
Above are the minimum configurations needed; you’ll further customize them in later examples.
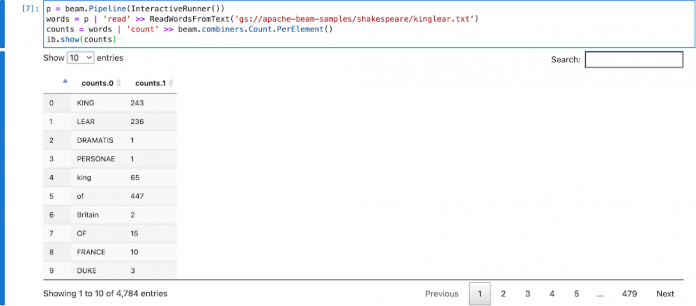
You can find the source code of the word count example here. Modify it with the interactive_flink_runner to build the pipeline in the notebook. The example uses gs://apache-beam-samples/shakespeare/kinglear.txt as the input file.
Inspecting the PCollection counts would implicitly start a Flink cluster, execute the pipeline, and render the result in the notebook.
Example 2 – Find out how many flights are delayed
This example reads more than 17 million records from a public BigQuery dataset, bigquery-samples.airline_ontime_data.flights, and counts how many flights have been delayed since 2010 for all the airlines.
On a normal InteractiveRunner running directly on a single notebook instance, it could take more than an hour for reading and processing due to the number of records (though the size of data is relatively small, ~ 1GB), and the pipeline can OOM or run out of disk space when the data is even bigger. With interactive Flink on notebook-managed clusters, you work with a higher capacity and performance (~ 4 mins for the example) while still being able to construct the pipeline step by step and inspect the results one by one within a notebook.
You need to have BigQuery service activated.
Configure a much bigger cluster with the options below3.
Whenever you inspect the result of a PCollection through ib.show() or ib.collect() in a notebook, Beam implicitly runs a fragment of the pipeline to compute the data. You can adjust the parallelism of the execution interactively.
With the above configurations, when you inspect data in the notebook, you are instructing Beam to implicitly start or reuse a Flink cluster on Google Cloud (Dataproc under the hood) with 40 VMs and run pipelines with parallelism set to 150.
You can include visualize_data=True when inspecting data through ib.show(). Binning the visualized data by their count, you can see that WN airline has the most delayed flights recorded in the dataset.
Example 3 – Run ML inference at scale interactively
The RunInference example classifies 50,000 image files (~280GB) from within the notebook.
The workload normally takes half a day for a single notebook instance or worker. With interactive Flink on notebook-managed clusters, it shows the result in ~1 minute. Looking at the Flink job dashboard, the actual inference only took a dozen seconds. The rest of the running time is overhead from staging the job, scheduling the tasks, writing the aggregated result to ib.options.cache_root, transferring the result back to the notebook, and rendering it in the browser.
Setup
For the RunInference example, you need to build a container image. You can find more information about building a container image from a notebook in this guide.
The extra Python dependencies needed for this example are:
The example uses the validation image set from ImageNet and the PyTorch pre-trained ImageNetV2 model. You can download similar dependencies4 or use your own image dataset and model. Make sure you copy the pre-trained model to the container and use its file path in the Beam pipeline.
Configure the pipeline options to use the custom container you build.
Build the pipeline
To run inference with a Beam pipeline, you need the following imports:
Then you can define processing logic for each step of the pipeline. You can use a mixture of DoFns and normal functions that yield or return and later incorporate them into the pipeline with different transforms.
Now define a few variables.
And build the pipeline with the above building blocks.
The pipeline reads a text file with 50,000 image file names in it. The Reshuffle is necessary to rebalance the image file names to all the workers before reading the image files. Without it, all 50,000 files will be read from a single task/thread/worker no matter how high the parallelism is.
Once read, each image will be classified into 1 of 1000 classes (e.g., a cat, a dog, a flower). The final aggregation counts how many images there are for each class.
In notebooks, Beam tries to cache the computed data of each PCollection that is assigned to a variable defined in the main module or watched by ib.watch({‘pcoll_name’: pcoll}). Here, to speed everything up, you only assign the final aggregation to a PCollection variable named counts as it’s the only data worth inspection.
To inspect the data, you can use either ib.show or ib.collect. If it’s the first time you inspect the data, a Flink cluster is implicitly started. For later inspections, computed PCollections do not incur executions. For inspections of data by newly appended transforms, the same cluster will be reused (unless instructed otherwise).
You can also inspect the cluster by running ib.clusters.describe(pipeline).
And you can follow the link in the output to the Flink dashboard where you can review finished jobs or future running jobs.
As you can see, the process took 1m45s to run inference for 50,000 images (~280GB).
You can further enrich the data if you know the mappings between classifications and their human-readable labels.
When inspecting the label_counts, the computed counts will be reused for the newly added transforms. After an aggregation, the output data size can be tiny compared with the input data. High parallelism does not help with processing small data and could introduce unnecessary overhead. You can interactively tune down the parallelism to inspect the result of processing only a handful of elements with the newly added transform.
Clean Up
Execute the code below to clean up clusters created by the notebook and avoid unintended charges.
Optionally, you can go to the Dataproc UI to manually manage your clusters.
Open Source Support
Apache Beam is open source software. The interactive features work with all IPython kernel-backed notebook runtimes. This also means the interactive FlinkRunner feature can be adapted to your own notebook and cluster setups.
For example, you can use Google Colab (a free alternative to Dataflow-hosted Beam Notebooks) connected with a local runtime (kernel) on your own workstation and then interactively submit jobs to a Flink cluster that you host and manage.
Set up Google Colab with local runtime
Set up a Flink cluster locally
To use your own Flink cluster, simply specify the necessary options:
If you use Beam built from source code (a dev version), you can configure a compatible container image.
Now you can run Beam pipelines interactively at scale on your own setup.
Compatibilities
Interactive Flink features are not patched back to older versions of (Interactive) Beam. Here is a compatibility table.
Beam Versions
Dataflow-hosted Beam Notebooks
Other notebook and cluster setups
<2.40.0
Not supported
Not supported
>=2.40.0,<2.43.0
Supported
Parallelism fixed to 1
>=2.43.0
Supported
Supported
There is also a cluster manager UI widget in the JupyterLab extension apache-beam-jupyterlab-sidepanel. Dataflow-hosted Beam Notebooks have it pre-installed. If you use your own JupyterLab setup, you can install it from either NPM or source code. It’s not supported in other notebook runtime environments such as Colab or classic Jupyter Notebooks.
Next Steps
Go to the Vertex AI workbench and get started using Dataflow-hosted Beam Notebooks! You can create, share, and collaborate on your notebooks with ease. And you have the flexibility to control who can access your notebook and what resources to use any time you want to make a change.
For the interactive Flink feature, check the public documentation for tips, caveats and FAQs when you run into issues.
Your feedback, suggestions, and open source contributions are welcomed.
Cloud BlogRead More