Are you looking to migrate a large amount of Hive ACID tables to BigQuery?
ACID enabled Hive tables support transactions that accept updates and delete DML operations. In this blog, we will explore migrating Hive ACID tables to BigQuery. The approach explored in this blog works for both compacted (major / minor) and non-compacted Hive tables. Let’s first understand the term ACID and how it works in Hive.
ACID stands for four traits of database transactions:
Atomicity (an operation either succeeds completely or fails, it does not leave partial data)
Consistency (once an application performs an operation the results of that operation are visible to it in every subsequent operation)
Isolation (an incomplete operation by one user does not cause unexpected side effects for other users)
Durability (once an operation is complete it will be preserved even in the face of machine or system failure)
Starting in Version 0.14, Hive supports all ACID properties which enables it to use transactions, create transactional tables, and run queries like Insert, Update, and Delete on tables.
Underlying the Hive ACID table, files are in the ORC ACID version. To support ACID features, Hive stores table data in a set of base files and all the insert, update, and delete operation data in delta files. At the read time, the reader merges both the base file and delta files to present the latest data. As operations modify the table, a lot of delta files are created and need to be compacted to maintain adequate performance. There are two types of compactions, minor and major.
Minor compaction takes a set of existing delta files and rewrites them to a single delta file per bucket.
Major compaction takes one or more delta files and the base file for the bucket and rewrites them into a new base file per bucket. Major compaction is more expensive but is more effective.
Organizations configure automatic compactions, but they also need to perform manual compactions when automated fails. If compaction is not performed for a long time after a failure, it results in a lot of small delta files. Running compaction on these large numbers of small delta files can become a very resource intensive operation and can run into failures as well.
Some of the issues with Hive ACID tables are:
NameNode capacity problems due to small delta files.
Table Locks during compaction.
Running major compactions on Hive ACID tables is a resource intensive operation.
Longer time taken for data replication to DR due to small files.
Benefits of migrating Hive ACIDs to BigQuery
Some of the benefits of migrating Hive ACID tables to BigQuery are:
Once data is loaded into managed BigQuery tables, BigQuery manages and optimizes the data stored in the internal storage and handles compaction. So there will not be any small file issue like we have in Hive ACID tables.
The locking issue is resolved here as BigQuery storage read API is gRPC based and is highly parallelized.
As ORC files are completely self-describing, there is no dependency on Hive Metastore DDL. BigQuery has an in-built schema inference feature that can infer the schema from an ORC file and supports schema evolution without any need for tools like Apache Spark to perform schema inference.
Hive ACID table structure and sample data
Here is the sample Hive ACID table “employee_trans” Schema
This sample ACID table “employee_trans” has 3 records.
For every insert, update and delete operation, small delta files are created. This is the underlying directory structure of the Hive ACID enabled table.
These ORC files in an ACID table are extended with several columns:
Steps to Migrate Hive ACID tables to BigQuery
Migrate underlying Hive table HDFS data
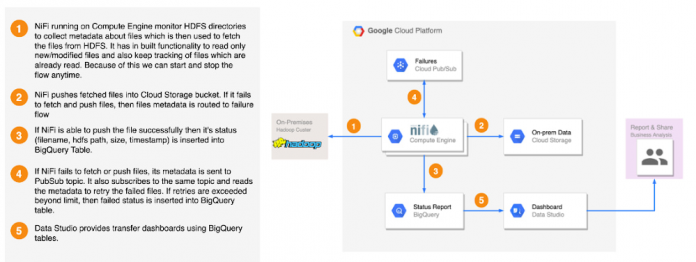
Copy the files present under employee_trans hdfs directory and stage in GCS. You can use either HDFS2GCS solution or Distcp. HDFS2GCS solution uses open source technologies to transfer data and provide several benefits like status reporting, error handling, fault tolerance, incremental/delta loading, rate throttling, start/stop, checksum validation, byte2byte comparison etc. Here is the high level architecture of the HDFS2GCS solution. Please refer to the public github URL HDFS2GCS to learn more about this tool.
The source location may contain extra files that we don’t necessarily want to copy. Here, we can use filters based on regular expressions to do things such as copying files with the .ORC extension only.
Load ACID Tables as-is to BigQuery
Once the underlying Hive acid table files are copied to GCS, use the BQ load tool to load data in BigQuery base table. This base table will have all the change events.
Data verification
Run “select *” on the base table to verify if all the changes are captured.
Note: Use of “select * …” is used for demonstration purposes and is not a stated best practice.
Load target BigQuery table
Use the below query to reflect the changes (updates and deletes) and load into the target table.
Once the data is loaded in target BigQuey table, you can perform validation using below steps:
a. Use the Data Validation Tool, or DVTfor short, to validate the Hive ACID table and the target BigQuery table. DVT provides an automated and repeatable solution to perform schema and validation tasks. This tool supports the following validations:
Column validation (count, sum, avg, min, max, group by)
Row validation (BQ, Hive, and Teradata only)
Schema validation
Custom Query validation
Ad hoc SQL exploration
b. If you have analytical HiveQLs running on this ACID table, translate them using the BigQuery SQL translation service and point to the target BigQuery table.
Hive DDL Migration (Optional)
Since ORC is self-contained, leverage BigQuery’s schema inference feature when loading.
There is no dependency to extract Hive DDLs from Metastore.
But if you have an organization-wide policy to pre-create datasets and tables before migration, this step will be useful and will be a good starting point.
a. Extract Hive ACID DDL dumps and translate them using BigQuery translation service to create equivalent BigQuery DDLs.
There is a Batch SQL translation service to bulk translate exported HQL (Hive Query Language) scripts from a source metadata bucket in Google Cloud Storage to BigQuery equivalent SQLs into a target GCS bucket.
You can also use BigQuery interactive SQL translator which is a live, real time SQL translation tool across multiple SQL dialects to translate a query like HQL dialect into a BigQuery Standard SQL query. This tool can reduce time and effort to migrate SQL workloads to BigQuery.
b. Create managed BigQuery tables using the translated DDLs.
Here is the screenshot of the translation service in the BigQuery console. Submit “Translate” to translate the HiveQLs and “Run” to execute the query. For creating tables from batch translated bulk sql queries, you can use Airflow BigQuery operator (BigQueryInsertJobOperator) to run multiple queries
After the DDLs are converted, copy the ORC files to GCS and perform ELT in BigQuery.
The pain points of Hive ACID tables are resolved when migrating to BigQuery. When you migrate the ACID tables to BigQuery, you can leverage BigQuery ML and GeoViz capabilities for real-time analytics. If you are interested in exploring more, please check out the additional resources section.
Additional Resources
Cloud BlogRead More